Running the Pipeline¶
The RECOVAR pipeline takes particle images and a mask, then computes the mean reconstruction, covariance, principal components, and embeddings.
Choose your workflow: CLI or GUI
This page has tabbed instructions for both the command line and the web GUI. Click the tab headers below each section to switch. Your choice is remembered across all pages. How to launch the GUI →
Submitting a pipeline job¶
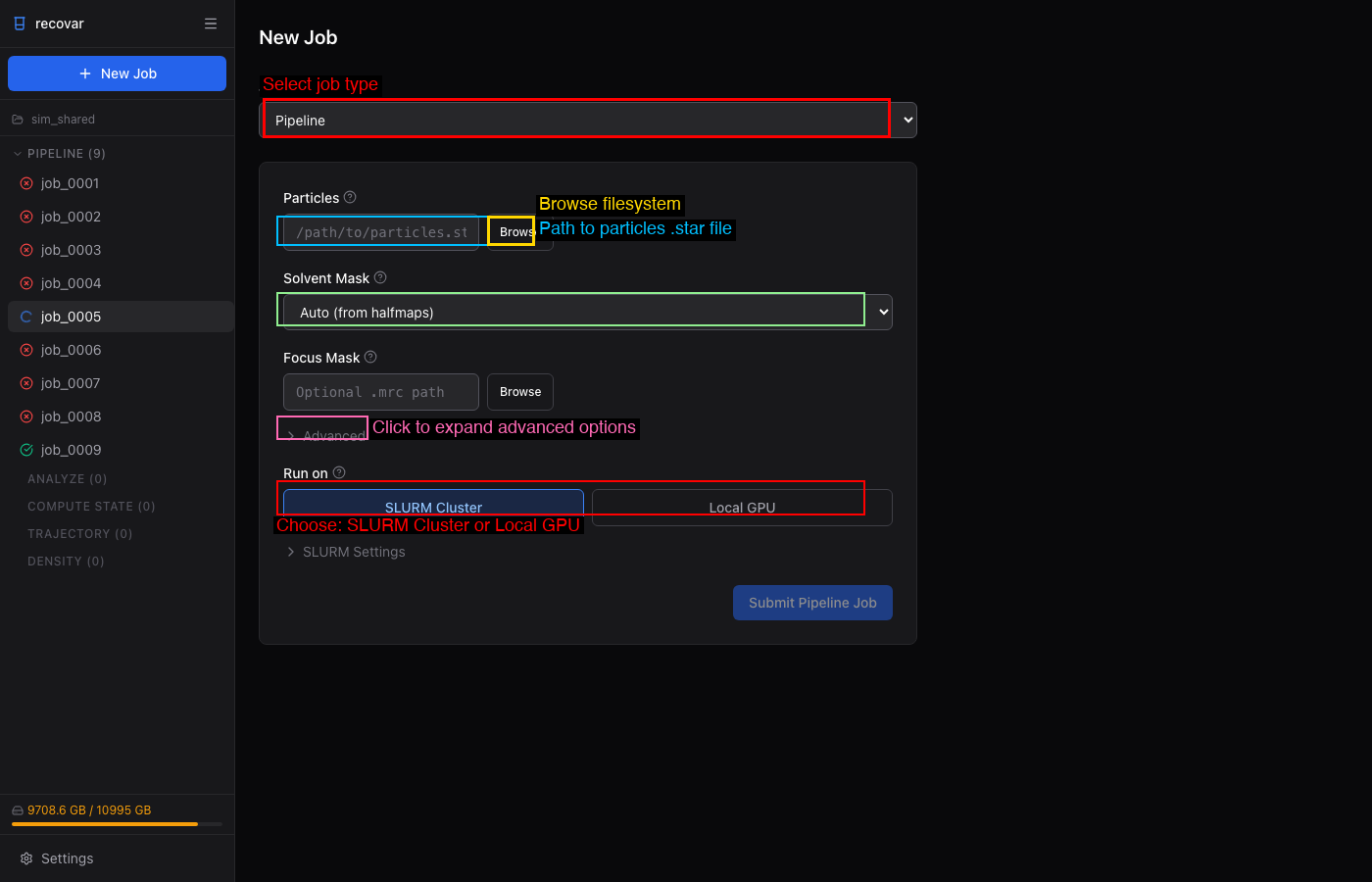
- Click + New Job in the sidebar
- Select Pipeline from the Job Type dropdown
- Browse to your particles file (
.star,.cs, or.mrcs) - Choose a solvent mask (Auto, Sphere, None, or custom
.mrc) - Choose SLURM Cluster or Local GPU under "Run on"
- Click Submit Pipeline Job
# Recommended: run inside a project
recovar init_project my_project
cd my_project
recovar pipeline particles.star --mask mask.mrc --project .
# cryoSPARC cs file
recovar pipeline particles.cs --mask mask.mrc --datadir /project/ --project .
# With downsampling
recovar pipeline particles.star --mask mask.mrc --downsample 128 --project .
# Standalone explicit output directory (still supported)
recovar pipeline particles.star -o output --mask mask.mrc
# Legacy pickle files
recovar pipeline particles.128.mrcs -o output \
--poses poses.pkl --ctf ctf.pkl --mask mask.mrc
Required arguments¶
| Argument | Description |
|---|---|
particles |
Input particles (.star, .cs, .mrcs, or .txt) |
-o, --outdir |
Output directory (optional in project mode) |
--mask |
Solvent mask (.mrc), or from_halfmaps, sphere, none |
Common options¶
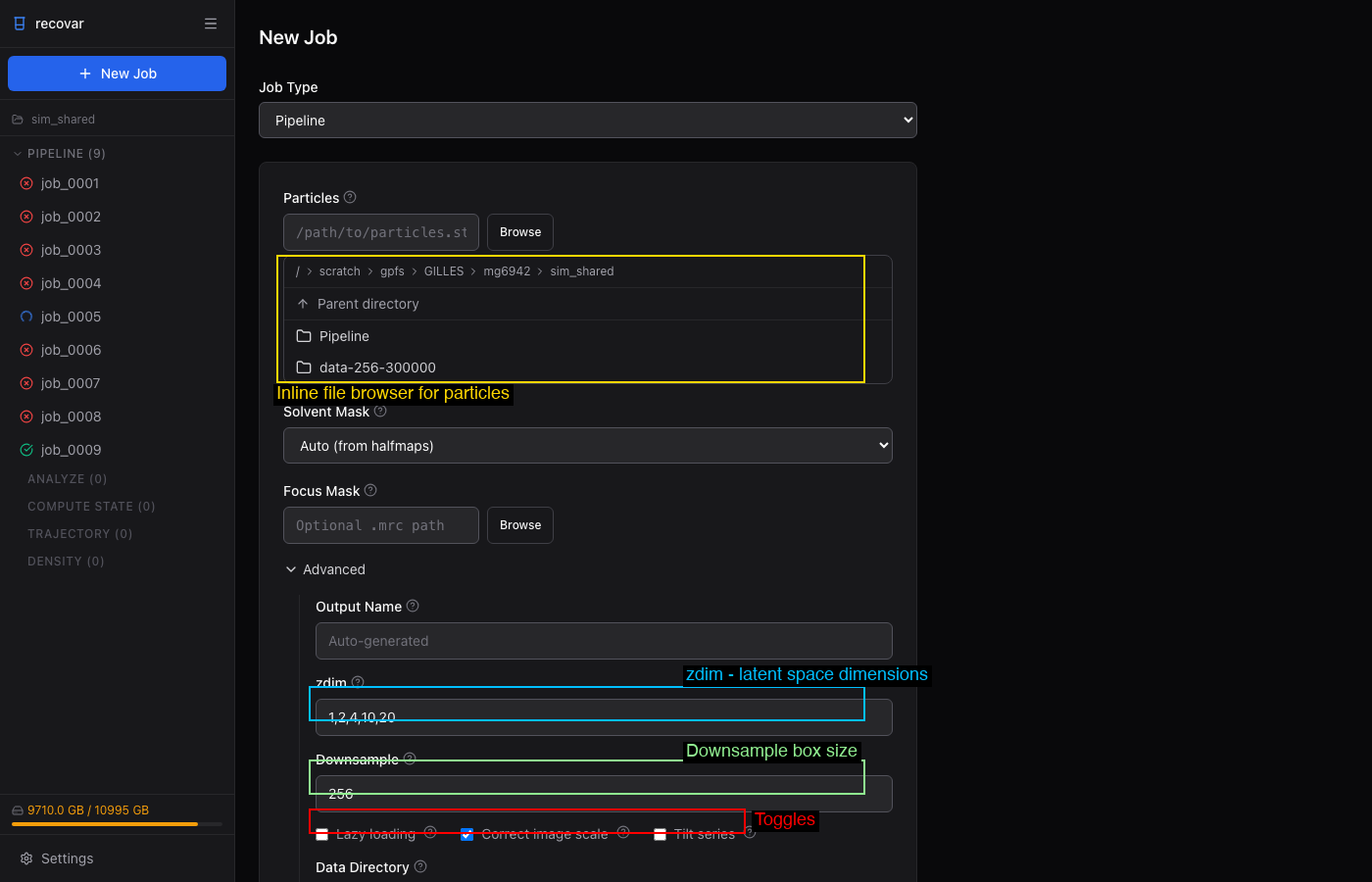
Expand the Advanced section in the job form to set:
- zdim -- PCA dimensions (default: 1,2,4,10,20)
- Downsample -- target box size (e.g., 128)
- Lazy loading -- for large datasets
- Correct image scale -- amplitude scaling correction
- Focus Mask -- browse to a custom focus mask
- Tilt series -- enable for cryo-ET data
Under Rarely Used: Poses, CTF, N Images, Halfsets.
| Flag | Default | Description |
|---|---|---|
--downsample D |
None | Downsample images to box size D (pre-downsamples to disk) |
--poses |
Auto | Poses file (.pkl). Auto-extracted from .star/.cs |
--ctf |
Auto | CTF file (.pkl). Auto-extracted from .star/.cs |
--focus-mask |
None | Focus mask for targeted heterogeneity |
--mask-dilate-iter |
0 | Dilate the mask by this many iterations |
--zdim |
1,2,4,10,20 | PCA dimensions for embedding |
--only-mean |
False | Only compute the mean (fast, for verifying setup) |
--correct-contrast |
False | Estimate and correct amplitude scaling |
--lazy |
False | Lazy loading for large datasets |
--multi-gpu |
False | Multi-GPU parallelization (experimental) |
Execution settings¶
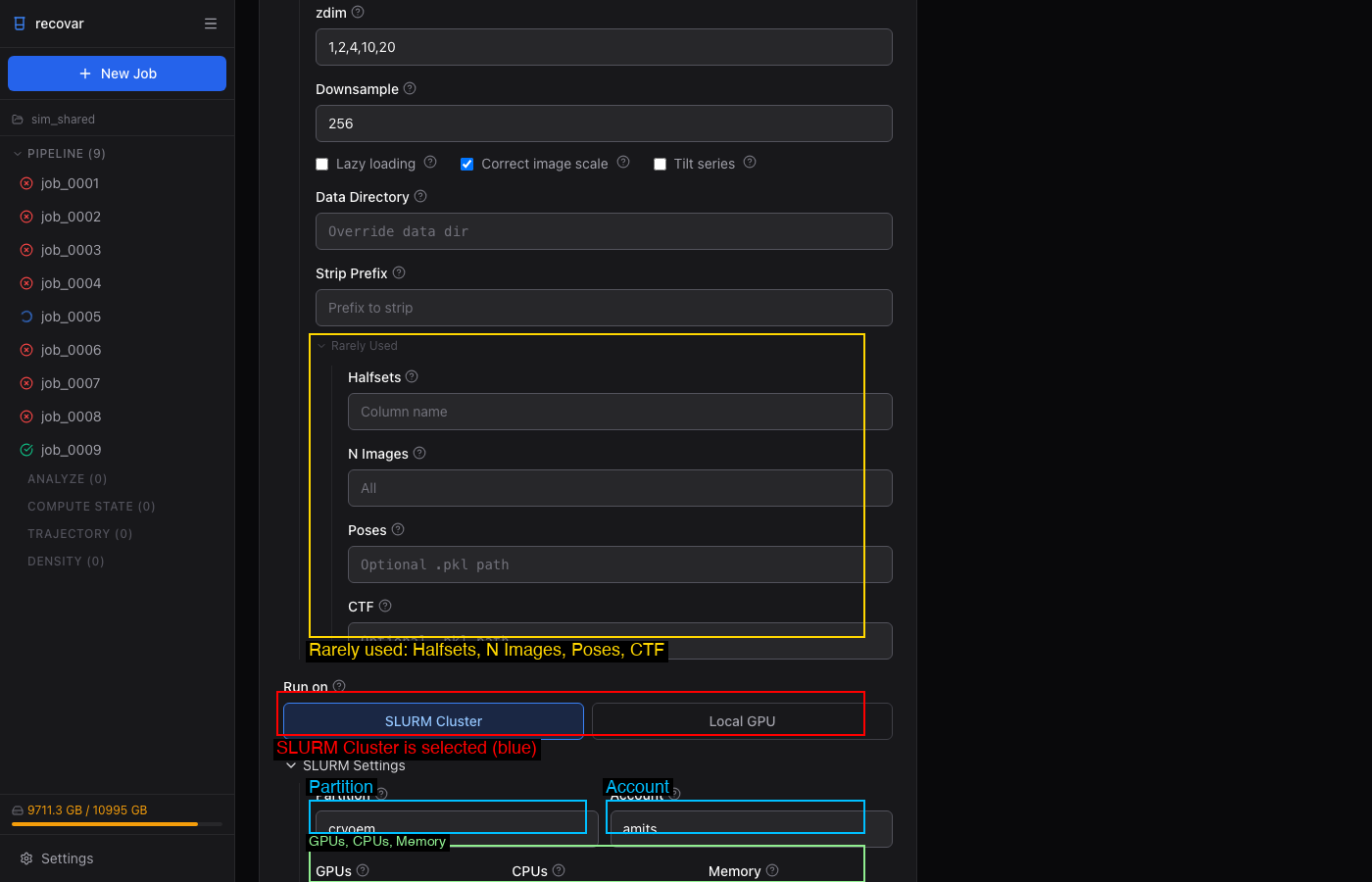
When submitting to SLURM (either from the GUI or CLI on a cluster), configure:
- Partition and Account — your cluster allocation
- GPUs, CPUs, Memory, Time limit
These can be saved as defaults in Settings (gear icon in sidebar) so you don't have to fill them in every time.
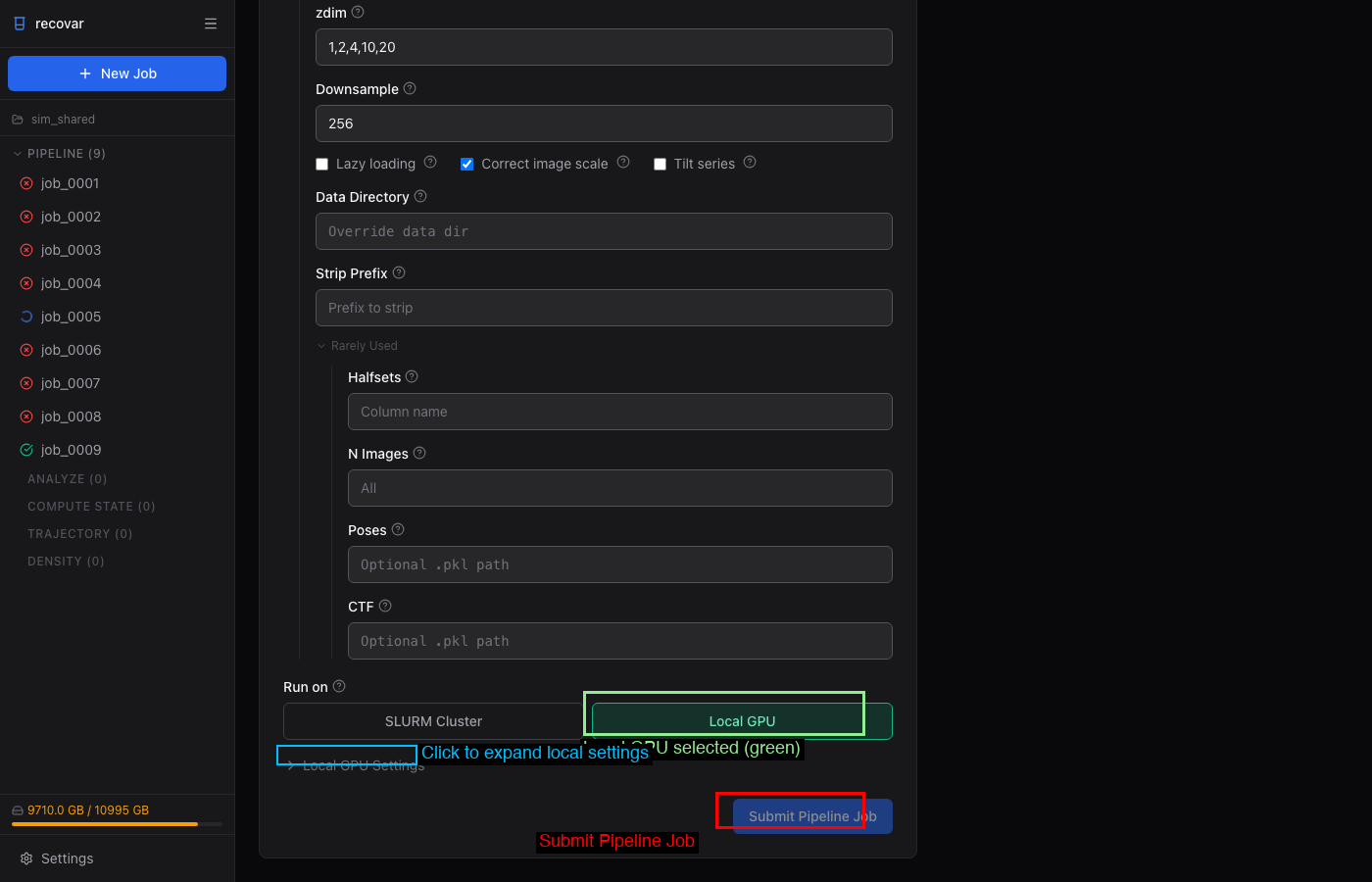
Run directly on the current machine's GPUs without SLURM:
- GPU picker — select specific GPUs or use all
- Setup command — e.g.,
module load cudatoolkit/12.8 - Environment variables — extra env vars for the job
Dataset loading options¶
| Flag | Default | Description |
|---|---|---|
--datadir |
None | Path prefix for resolving relative image paths |
--strip-prefix |
None | Strip prefix from image paths |
--ind |
None | Filter to specific image indices (.pkl) |
--particle-ind |
None | Filter particles by indices (cryo-ET only, .pkl) |
--n-images |
All | Number of images to use |
--halfsets |
None | Pre-computed half-set split (.pkl) |
--padding |
0 | Real-space padding |
--uninvert-data |
automatic | Data sign inversion: true, false, or automatic |
Advanced options¶
| Flag | Default | Description |
|---|---|---|
--noise-model |
radial | Noise model: radial or white |
--mean-fn |
triangular | Mean function: triangular or triangular_reg |
--gpu-gb |
All | GPU memory limit in GB |
--n-gpus |
All | Number of GPUs to use |
--keep-intermediate |
False | Save intermediate results |
--accept-cpu |
False | Allow running without GPU |
--ignore-zero-frequency |
False | Useful if images are normalized to zero mean |
--low-memory-option |
False | Lower memory for covariance estimation |
--very-low-memory-option |
False | Lowest memory for covariance estimation |
--premultiplied-ctf |
False | Input images have pre-multiplied CTF |
Multi-GPU (experimental)¶
Multi-GPU support parallelizes the covariance estimation step across GPUs. This is the most expensive step of the pipeline, so multi-GPU can significantly reduce total runtime for large datasets.
# Use all available GPUs
recovar pipeline particles.star -o output --mask mask.mrc --multi-gpu
# Use specific number of GPUs
recovar pipeline particles.star -o output --mask mask.mrc --multi-gpu --n-gpus 4
Work in progress
Multi-GPU is experimental. It parallelizes covariance estimation only — the mean reconstruction and embedding steps still run on a single GPU. If you run into issues, drop --multi-gpu and the pipeline will run normally on one GPU.
GPU memory and device selection¶
# Limit memory per GPU (useful on shared machines)
recovar pipeline particles.star -o output --mask mask.mrc --gpu-gb 8
# Select specific GPUs by ID
CUDA_VISIBLE_DEVICES=0,2 recovar pipeline particles.star -o output --mask mask.mrc --multi-gpu
# Disable JAX memory preallocation (useful on shared machines)
XLA_PYTHON_CLIENT_PREALLOCATE=false recovar pipeline ...
Cryo-ET options¶
| Flag | Default | Description |
|---|---|---|
--tilt-series |
False | Use tilt-series data |
--tilt-series-ctf |
Auto | CTF model: cryoem, relion5, warp |
--dose-per-tilt |
From file | Dose per tilt |
--angle-per-tilt |
From file | Angle per tilt |
--ntilts |
All | Number of tilts per series |
See Cryo-ET for details.
Output structure¶
output/
job.json # Job metadata (version, timing, parameters)
command.txt # Command that was run
run.log # Full log
README.txt # Human-readable output summary
downsampled/ # Pre-downsampled images (if --downsample used)
particles.128.mrcs
particles.128.star
model/ # Internal model data
params.pkl # Pipeline parameters
zdim_4/ # Per-zdim embedding directories
latent_coords.npy # Latent coordinates for zdim=4
zdim_10/
latent_coords.npy
...
output/
volumes/
mean.mrc # Mean reconstruction
mean_filt.mrc # Filtered mean
mean_half1_unfil.mrc # Unfiltered half-map 1
mean_half2_unfil.mrc # Unfiltered half-map 2
mask.mrc # Solvent mask used
dilated_mask.mrc # Dilated mask
plots/ # Diagnostic plots
analysis_*/ # Results per zdim (after running analyze)
When using the project system (--project), pipeline output is placed into auto-numbered directories like Pipeline/job_0001/. The numbered directories stay stable on disk, while RECOVAR records human-readable job names in project metadata for the CLI and GUI.
Viewing results¶
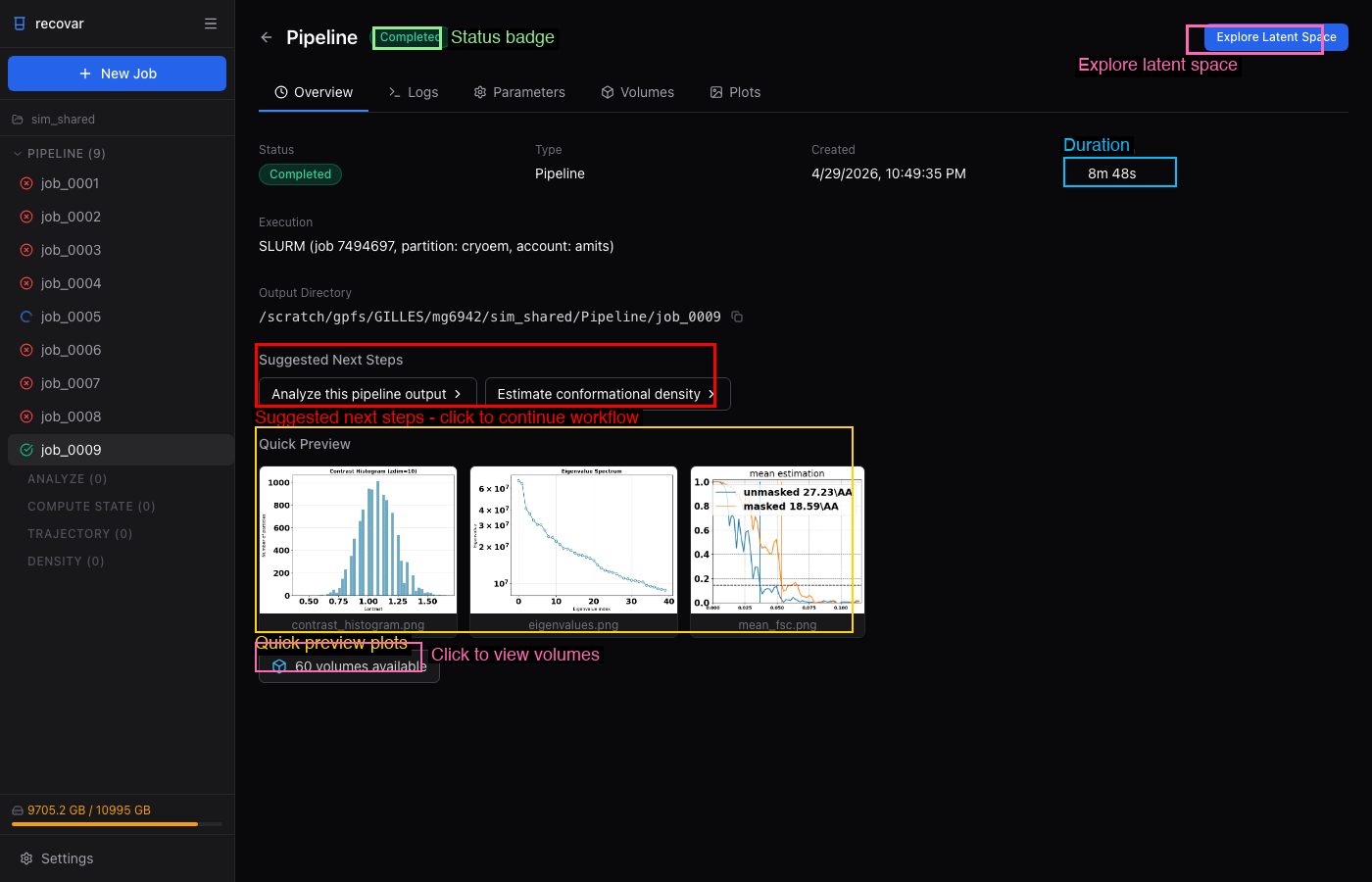
After the pipeline completes, the job detail page shows:
- Quick Preview -- contrast histogram, eigenvalue spectrum, mean FSC
- Volumes tab -- browse all output volumes (mean, eigenvolumes, variance maps)
- Plots tab -- all diagnostic plots
- Suggested Next Steps -- one-click links to run Analyze or Density Estimation
See the Tutorial for a full worked example with real pipeline output and plots on EMPIAR-10076 (50S ribosome, 131k particles).
Tips¶
Recommended starting parameters
- Downsample: 128 for speed, 256 for quality
- zdim: Start with
1,2,4,10,20(default). For publication, also try--zdim 40 - n-images: Use 10000-50000 for initial exploration, all images for final run
- --correct-contrast: Always enable this unless you know your data is already contrast-corrected
Quick setup check
Use --only-mean for a fast run that only computes the mean reconstruction. This verifies your data, mask, and CTF are correct before committing to a full run.
Large datasets
For datasets > 500k particles, use --lazy for lazy loading and --downsample 128 for speed. Consider --n-images 100000 for initial exploration.
Memory
If you run out of GPU memory, try --low-memory-option or --very-low-memory-option. You can also limit memory with --gpu-gb 8.